



The ENCODE (Encyclopedia of DNA Elements) Consortium is an international collaboration of research groups funded by the National Human Genome Research Institute (NHGRI). Its goal is to build a comprehensive parts list of functional elements in the human genome. The catalog includes genes (protein-coding and non-protein coding), transcribed regions, and regulatory elements, as well as information about the tissues, cell types and conditions where they are found to be active.
The ENCODE web portal allows researchers to perform queries using various parameters such as assay, biosample, and genomic annotations. A typical researcher workflow involves searching for data on the ENCODE portal, downloading relevant datasets individually, keeping track of all data that are downloaded, running various analyses on the data, creating results, and eventually publishing conclusions.
While online access to ENCODE data is a great boon to research, subsequent steps can become cumbersome. Each data file URL returned by a query must be downloaded individually, and data comes without associated metadata or context. Researchers must manually save queries if they wish to record data provenance. The provenance of intermediate datasets and analysis results is often lost unless the researcher diligently captures them. There is no way for a researcher to validate that a copy of an ENCODE dataset has not been corrupted, other than to download the data again.
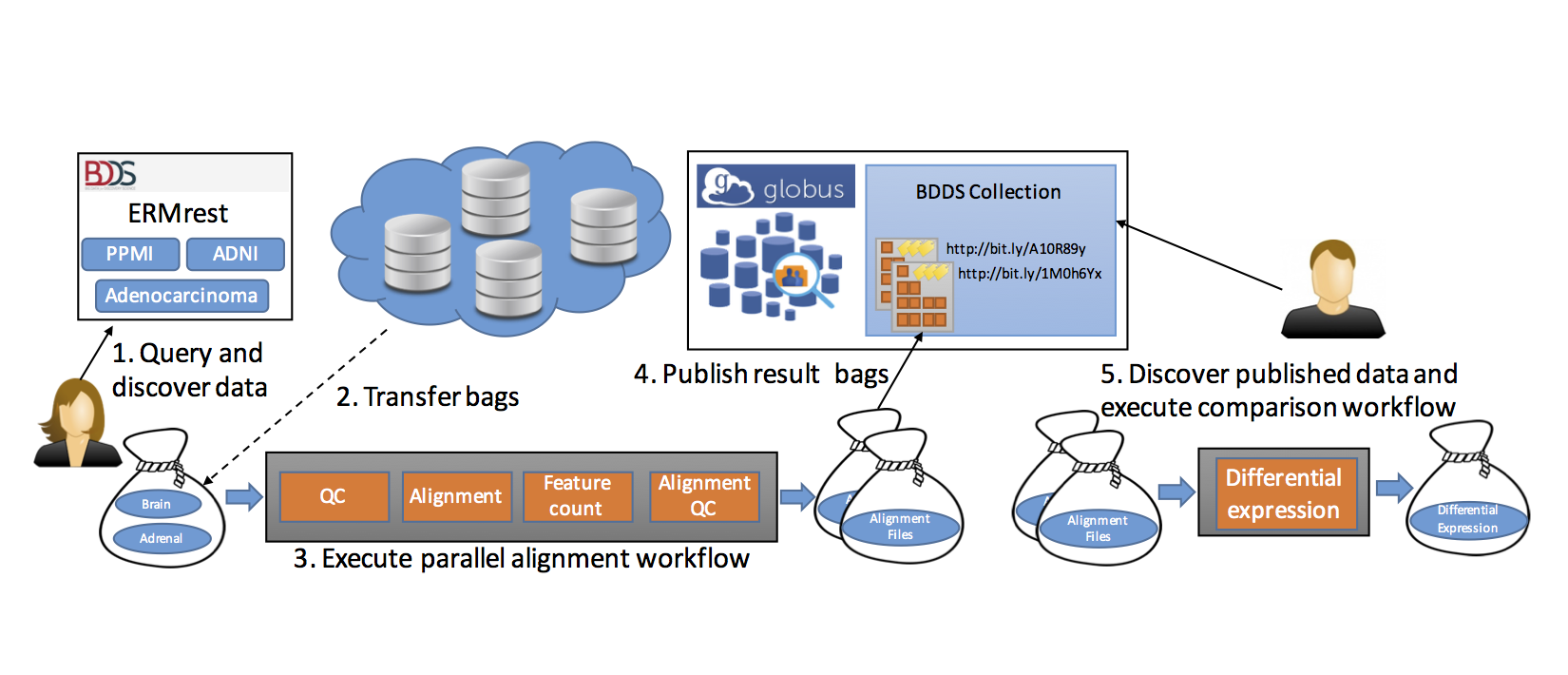
We describe in the following how our tools can be used to realize the end-to-end scenario shown in Figure below.
We used BDBag, Minid, and Globus tools to create a simple portal that allows the researcher to access the results of an ENCODE query, plus associated metadata and checksums, as a BDBag. Figure shows this portal in action. The researcher enters an ENCODE query in REST format or uploads an ENCODE metadata file that describes a collection of datasets. The query shown in Figure identifies data resulting from RNA- Seq experiments on stem cells. (As of August 2016, the result comprises 13 datasets with 144 FastQ, BAM, .bigWig, and other files, totaling 655 GB of the 193 TB in ENCODE.)
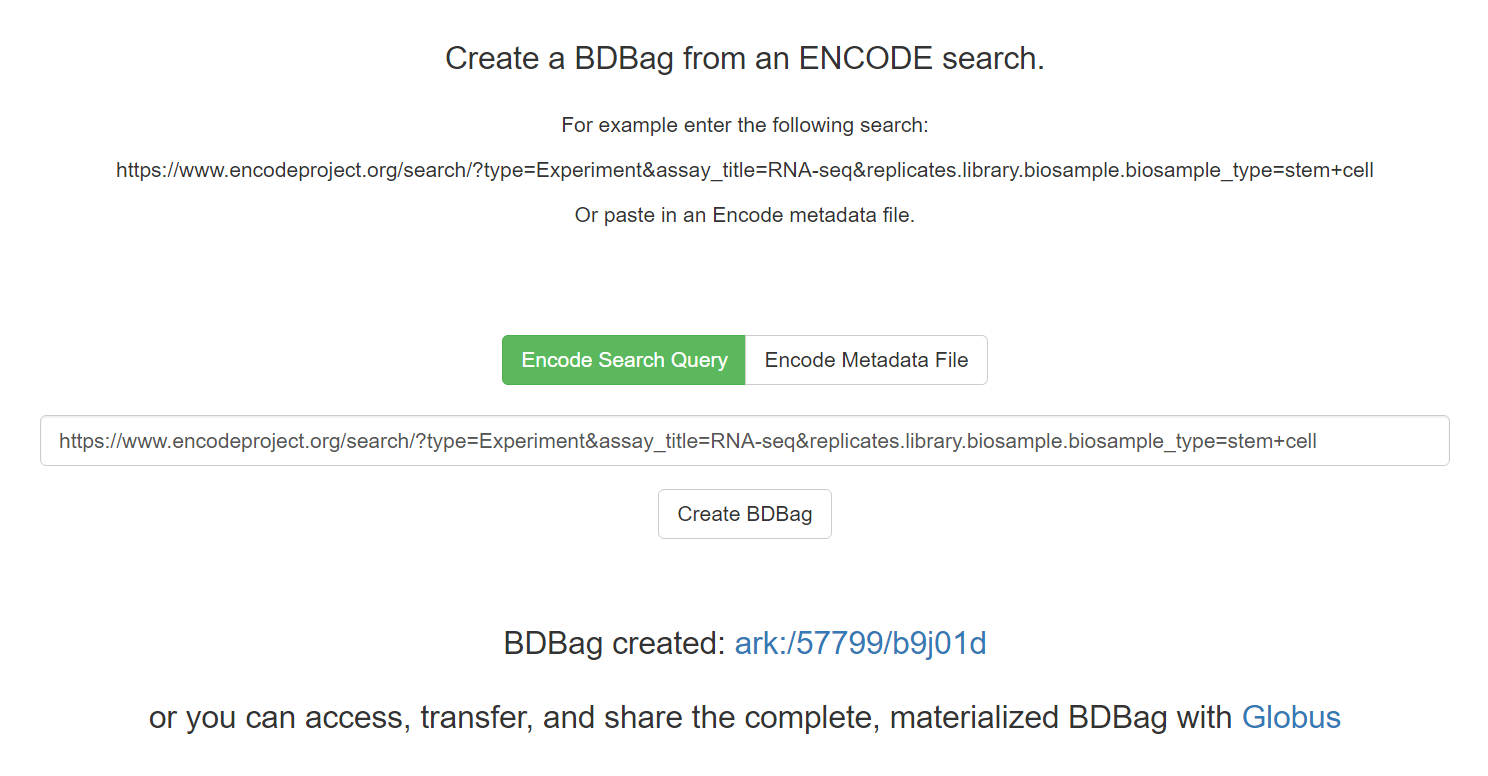
The researcher can then select the “Create BDBag” button to trigger the creation of a 100 KB BDBag, stored on Amazon S3 storage, that encapsulates references to the files in question, metadata associated with those files, and the checksums required to validate the files and metadata.
As shown in Figure below, the user is provided with a Minid for the BDBag, ark:/57799/b9j01d.

This identifier leads to a landing page similar to that shown in Figure, from which the user can download the BDBag. The BDBag is stored in Amazon S3 and can therefore be permanently referenced for purposes of sharing, reproducibility, or validation. The BDBag materialization tool can then be used to fetch each file from the URL provided in the fetch.txt file. Given the Minid for a bag, a user may copy the bag to a desired storage location and use the BDBag tools to retrieve remote files from the ENCODE repository. The default practice of making client-driven HTTP requests can be time consuming and is subject to failures. For example, we found that instantiating the 655 GB test bag of remote ENCODE data via HTTP took 36 hours and two restarts. The BDBag tools provides an effective alternative: if the remote data is located on a Globus endpoint, reliable and high performance Globus transfer methods can be used that improve performance and automatically restart interrupted transfers. Thus, for example, we were able to instantiate the same BDBag from Petrel, a storage repository at Argonne National Laboratory, to a computer at the University of Chicago Research Computing Center in 8.5 minutes, and from Amazon S3 to the same computer in 14 minutes. If BDBag elements are referenced using Minids, the BDBag tools have the option of preferring Globus URLs if they are provided. The checksumming of bag contents ensures that we have the correct data, regardless of source.
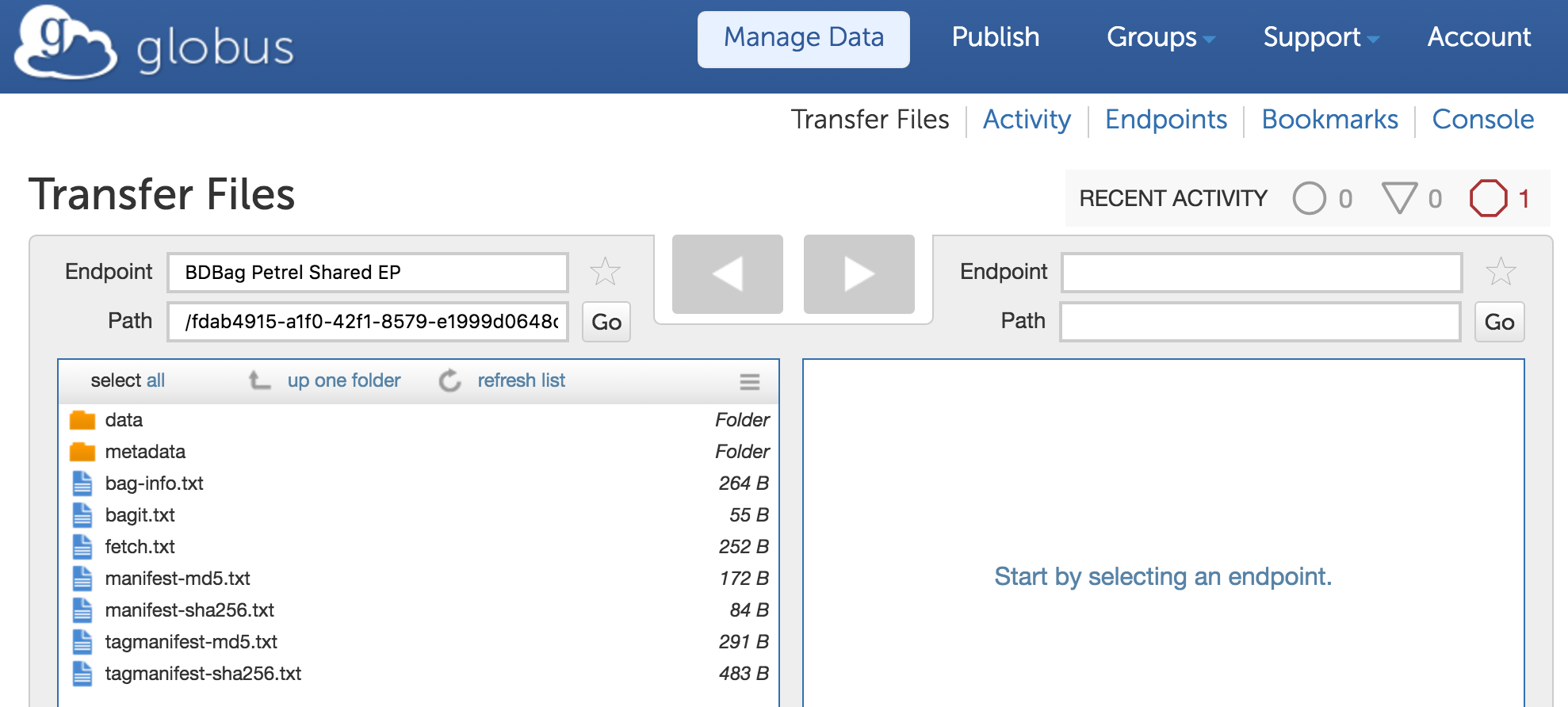
Having enabled direct, programmatic access to ENCODE query results plus associated metadata, we next want to automate the analysis of such results. To this end, we use the Galaxy-based Globus Genomics extended to resolve a BDBag via a Minid, accept BDBags as input, retrieve the referenced datasets, and run analytical pipelines at scale on cloud resources: Steps 2 and 3 in Figure below.
The analysis is performed on data from a molecular biology technique called DNase I hypersensitive sites sequencing, or DNase-seq. Such experiments generate a large volume of data that must be processed by downstream bioinformatics analysis. Currently available ENCODE data for our analysis protocol comprise 3–18 GB of compressed sequence. Depending on input sequence size, analysis can require 3–12 CPU hours per sample on a AWS node with 32 CPU cores. The final output file is relatively small compared to the input, but intermediate data is about 10 times the input size. The resulting output for each individual patient sample is encapsulated in a BDBag—containing a collection of candidate DNA footprints. All individual samples from the same cell line are merged and then filtered by intersecting against a database for known transcription factor binding sites in the reference genome. This analysis step takes one CPU hour and produces 50–100GB of output, depending on the cell type. The final BDBags are assigned a Minid. We also publish these BDBags into the BDDS Publication service (Step 4), so that other researchers can discover and access the results for other analyses (Steps 5 and 6).