
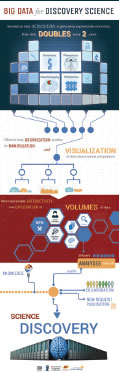



Biomedical data acquisition is generating exponentially more data: thousands of whole-genome sequences (WGS) are now available; brain data is doubling every two years. Analyses of Big Data, genomic or otherwise, presents qualitatively new challenges as well as opportunities. Please see abstract from American Society of Human Genetics Annual Meeting 2015. Among the challenges is a proliferation in ways analyses can fail, due largely to the increasing length and complexity of processing pipelines. Anomalies in input data, runtime resource exhaustion or unavailability of nodes in a distributed computation can all cause pipeline hiccups that are not necessarily obvious in the output. Flaws that can taint results may persist undetected in complex pipelines, a danger amplified by the fact that research is often concurrent with the development of the software on which it depends. On the positive side, the huge sample sizes increase statistical power, which in turn can motivate entirely new analyses.
BDDS is developing a fully automated tool for performing quality controls on big data. BDQC identifies anomalous files among large collections of files that are a priori assumed to be equivalent in structure and content. It was motivated by the realization that when running complex pipelines on large data sets, errors creep in for a variety of reasons, mostly related to the (unpredictable) runtime environment. BDQC is intended to 1) validate primary input data to pipelines; 2) validate output (or intermediate stages of) data processing pipelines; 3) discover potentially "interesting" outliers. Although it was developed in the context of genomics research, it is expressly not tied to a specific knowledge domain. It can be customized (via the plugin mechanism) for specific domain needs.
BDQC studies a collection of files that represent an equivalence class (e.g., input data, analysis results at a given stage in a pipeline, or the final results of the pipeline). For each file, it computes a series of automatically determined metrics and stores the result as a JSON object in associated *.bdqc files. This work is done using a robust Python3 framework that can be extended using plugins. These plugins embody a series of generic, domain-blind tests, as well as a series of domain-specific tests. These test results can then be analyzed to identify components (classes) and outliers.
The project is hosted on GitHub.