



The Big Data for Discovery Science Center (BDDS) is a unique effort focused on the user experience with big data. How are big data organized, managed, and stored? How are big data processed and distributed either to local or remote computing resources or to colleagues or to geographically distributed archives? How can a user focused on biology interact with vast collections and distant computers and storage systems to explore, interact and understand what the data mean and to derive knowledge from them? Tools that are not only enabling but intuitive and adaptive will be created to directly answer these needs. We have assembled a team comprised of leaders in computer science, neuroscience, genetics and knowledge discovery from the University of Southern California, the University of Chicago, the University of Michigan, and the University of Washington. We have a mature and competent staff of software developers, testers and applied scientists. And most importantly, we have experience in the challenges of big data and have developed solutions to portions of this significant and pressing big data need in biological research. Our center, entitled Big Data for Discovery Science will lead a new paradigm for interacting with large biomedical data types and scales - from ‘omes’ to ‘organs’.
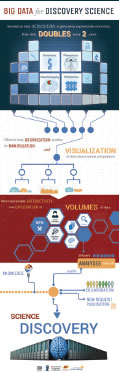
We have now entered an era of big data. Modern biomedical data collection, from genes to cells to systems, is generating exponentially more data due to increases in the speed and resolution of data acquisition methods. The thirst for still more data on fundamental biological processes, from the levels of “omics” to organs, then drives further technological innovations. This big data arises from varying sources and topologies - single large data sets from the efforts of individual laboratories, a collection of more modest studies across common study protocols, or amassed as a heterogeneous collection of research data. How and by what means data are processed and analyzed is an equally important element. New, user-focused tools are required to organize, manipulate, and mine largescale data, thereby maximizing the knowledge available for critical research questions. To inform a new generation of data scientists, dedicated training programs and workshops are needed on big data best-practices and potential. Our BD2K center team is comprised of leading biology and computer scientists, with expertise in large-scale biomedical data, and knowledge of the present challenges and promise of big datamechanisms in place.
Big-Data-driven decision-making is now broadly acknowledged and accepted as a new fact of life for many. The emergence of increasing data size creates exciting opportunities for a new mode of scientific discovery in which alternative hypotheses are developed, tested, and evolved against large existing data collections, rather than by generating data for the sole purpose of validating a predetermined hypothesis. First identified over a decade ago by Jim Gray, this data-driven research paradigm, which he called the fourth paradigm (with experimental, observational and model driven science being the previous three paradigms), has been codified in a scientific approach that has come to be called “discovery science” (Hey, Tansley et al. 2009). The hallmark of discovery science is that hypothesis are rapidly generated and tested against data that has been collected because of its potential to answer a range of questions. In astronomy, for example, digital sky surveys, which systematically image large fractions of the sky, allow astronomers to ask far-reaching questions via computer analysis of millions of stars and galaxies, rather than by personal observations of a few stars as in the past. A single survey can thus enable hundreds or even thousands of publications. In biomedicine, we see similar approaches, e.g. the creation of GWAS surveys that focus on data collection in the absence of specific hypothesis.
Many ongoing NIH-funded data collection activities represent biomedical approaches for big data collection, aggregation, and sharing. The means by which such big data collections and repositories get created and populated is depicted in Figure 2. In addition to data gathered by individual laboratories, this represents a variety of topologies for big data aggregation and exchange from singularly large data sets, through multi-site consortia, centralized curated archives, meta/ derived results, and federated databases. From these, collaborators may, in principle, access raw or summary information for use in novel processing, confirmatory analyses, or in combination as a mega-analysis across subjects/specimen, age groups, etc.
However, for the biomedical sciences, there is currently a wide gap between the potential for these big data resources and their realization as resources for discovery. Heterogeneity, scale, timeliness, complexity, incongruence, missingness, and privacy problems with biomedical research data can impede progress at all phases of the knowledge extraction workflow. Biomedical experimentation, while attempting to be empirically precise, is frequently comprised of highly unstructured and poorly described components such as file structures, formats, and results. The problems begin during data acquisition, when decisions are required, currently in an ad hoc manner, about what data to keep and what to discard, and how to store what we keep reliably with the right metadata. The value of biomedical data in one domain is enriched when it can be linked with other data from an entirely different domain, thus data integration is a major creator of value. One consequence of this is that users find data must often be assembled by combining data from different sources, locations, and types. Data analysis, organization, retrieval, and modeling are other foundational challenges. Data analysis for such data is a clear bottleneck in many applications, both due to lack of scalability of the underlying algorithms, and due to the complexity of the data that needs to be analyzed. Finally, presentation of the results and its interpretation by non-technical domain experts is crucial to extracting actionable knowledge.
How Big is “Big Data”?
Size is a relative term when it comes to data. Biomedical data frequently comes in a variety of forms with each generating differing types and amounts of information about biological structure and/or function. Moreoever, in vivo biomedical datatypes are often not unimodal but, rather, remarkably diverse - examining brain form, function, and connectivity, and rapidly improving its ability to resolve finer spatio-temporal scales. As biomedicial technologies have improved, been extended, or made faster, so too has the amount of data which can be obtained. Once these improved methodologies have proven robust and dependable, with analytic methods available with which to use them, researchers have been quick to adopt them – doubling or tripling the amount of data they can then gather per subject by doing so (Van Horn and Toga 2013). Individually, the data sets from a single study themselves may not pose major difficulties for processing and analysis using exising algorithms and statistical methods. However, as data sets are amassed into large-scale databases, considerable challenges emerge. Such data will only grow over time and what “big” data is collected today will tomorrow seem “cute” by contrast. Along with the increasing interest in gathering study data from across the lifespan or focused on specific patient groups, and comprehensive phenomic meta-data on each study participant, wrangling, let alone interpreting, tomorrow’s biomedical data will not be for the faint of heart. Thus, it is safest to say that “Big Data” is not the data that we have now but the data that we will collect in the future. How we prepare for that future will make all the difference in our ability to leverage big data into new discoveries, knowledge, treatments, and cures.